Medicine in the 21st Century is becoming more and more personalized. So there is a focus on developing medicines that are tailored toward individual aspects of each patient instead of trying to treat everybody in the same way. Personalized medicine depends on testing a patient for some aspect of their disease, called a biomarker.
A biomarker can be any molecule whose existence or absence is measurable and which gives information about the state of health. In particular, a biomarker differentiates tumor from healthy tissue. While pathogen-related diseases are easy target for drugs and immune system because the pathogen is easily recognizable, microscopic differences (biomarkers) distinguish tumor cells from normal ones. Therefore, the ability to spot biomarkers is essential to design the best therapy for cancer patients.
A biomarker test is performed in a pathology laboratory on a blood or tissue sample from the patient and the test result can be used to predict whether the patient can benefit from a particular medicine. There are over 100 personalized medicines now approved for use, and 50% of new drugs that are now in development are linked to biomarkers, so that number will continue to grow. In oncology, biomarkers often are related to alterations or mutations of genes that cause cancers to grow and spread. There are over 1000 ongoing clinical trials for cancer drugs that are linked to gene-based biomarkers, and 50 biomarker-linked therapies are already approved for clinical use. The presence of certain gene alterations or mutations in cancer can guide the use of targeted therapies whose mechanism of action is directly related to those alterations. There is growing evidence that a given gene mutation may be present in multiple different tumor types and we are starting to see some drugs being approved across cancer types, so these have pan-cancer or tumor-agnostic indication for specific gene mutations or alterations.
On the other hand, we also know that within a specific type of cancer, different patients have different gene mutations that cause the tumor to grow and spread. These different mutations are linked to different drug treatments. So, for a single tumor type, there are now multiple gene-directed therapies that have been approved. But while the development of these new drugs is greatly benefiting cancer patients, many times there is limited biopsy tissue available to test for all these different mutations and this is a growing challenge for oncologists and pathologists. For example, biopsies from lung cancer can be very small and have limited amount of tumor for testing the several, potentially-mutated genes. The preferred strategy is to screen first for the most frequently mutated ones, and then to screen the others, one by one, until (hopefully) a positive result is found. If the biopsy sample is not sufficient, this could limit the physician ability to offer some therapeutic options to the patient. That’s why biotech companies are now developing tools able to test all biomarkers at once (called multiplex tests). These tests are based on DNA sequencing and are called Comprehensive Genomic Profiling (CGP) tests. Some of the biomarkers that are measured by CGP are Micro-Satellite Instability (MSI), Tumor Mutational Burden (TMB) and fusion genes.
Microsatellite Instability
Micro-satellites are short repeats of nucleotides (usually 1 to 6 nucleotides repeats; the most common is the dinucleotide GT: GTGTGTGT…). These repetitive structures are fragile and prone to breakage. If the DNA repair mechanism does not function properly, mutations can accumulate at these sites, causing genomic instability. This leads to high mutations rate, which predisposes to tumors.
For example, the drug Pembrolizumab has a pan-cancer or tumor-type agnostic approval for use in “MSI-high” cancers (so cancers that have high mutations rate at MSIs).
Tumor Mutational Burden
TMB is measured by assessing the total number of mutations across a large expanse of coding genome. Cancer cells with high TMB are believed to express more antigens, which are recognized by immune cells. Scientists proved that “TMB-high” is a novel important biomarker for tumors, because these tumors provide better response to immunotherapy.
Despite apparent similarity, MSI and TMB are not the same thing and indeed their features do not overlap in tumors: there can be tumors with high mutation rate at micro-satellites (which is a small portion of the genome), but TMB-low, and vice versa, when a tumor has high mutational rate across the genome (TMB-high), but specifically not at micro-satellites (MSI-low).
Fusion genes
CGP provides information also about fusion genes: they are present in 15% of metastatic cancers and form when a piece of chromosome breaks and migrates to another location in the genome (often to a different chromosome). A gene present on the translocating DNA, could merge with another gene in the new location, possibly producing a mutant protein, which becomes a biomarker of the tumor. Detection of fusion genes is complex because there can be several fusion partners (even 60) for a single translocating gene. Because fusion partners affect sensitivity to drugs, it is important to identify the specific partners involved in the fusion.
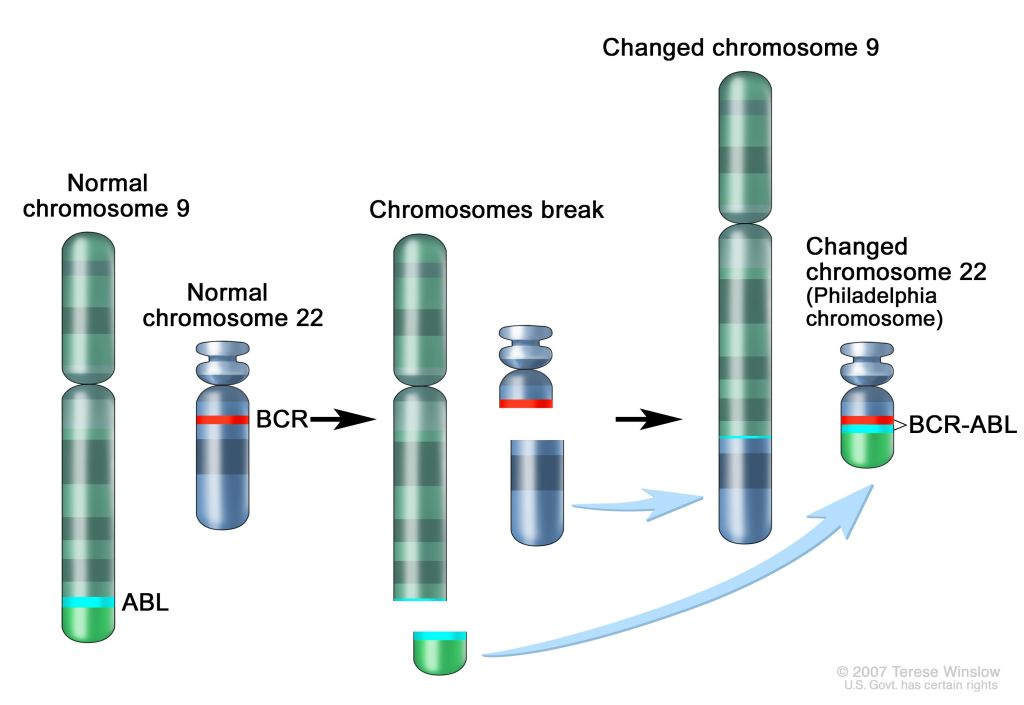
One bright example of how important is to identify fusion genes is represented by the case of chronic myeloid leukemia. In this type of blood cancer, the short bottom piece of chromosome 9 and the short top piece of chromosome 22 attach to one another. This translocation creates a longer chromosome 9 and a shorter chromosome 22 (see the figure below). The shorter chromosome 22 is called the Philadelphia chromosome. As a result of this translation, the ABL gene from chromosome 9 and the BCR gene from chromosome 22 join together and form the BCR-ABL fusion gene, the biomarker of this type of cancer. Against this biomarker, it was developed a monoclonal antibody called imatinib (followed then by others), which inhibits the progression of 75% of cancers expressing BCR-ABL fusion gene.